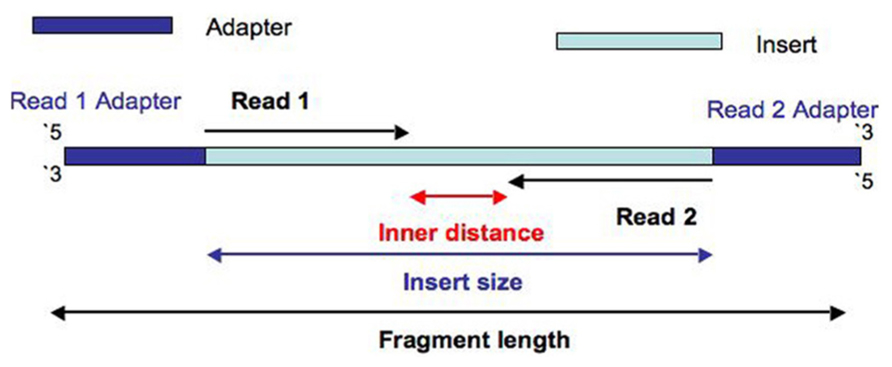
  Cell Multiplexing library contains Illumina Nextera Read 1 and Nextera Read 2 adapter sequences. Up to 384 uniquely indexed samples may be pooled and sequenced together. Nextera Read 1 and TruSeq Read 2 adapter sequences: CSP.png. (For blood and saliva, see the reference guide).Ä¡6s rRNA Sequencing, Amplicon Sequencing, De Novo Sequencing, Shotgun Sequencing, Whole-Genome SequencingĪmplicon Sequencing, De Novo Sequencing, Shotgun Sequencing, Whole-Genome Sequencing ~3-4 hours (from DNA extraction to normalized library)įast library prep optimized for research on small genomes, PCR amplicons, and plasmids.Ī fast, flexible workflow for a wide range of research applications and sample types, from human to microbial whole-genome sequencing and more. Nextera XT Index Kit v2 adapter Trim bases from sequences2, 3 Trim bases. ~5.5 hours from DNA extraction to normalized library. Illumina Qualified Methods are available on a range of automation platforms through our partners. Nextera transposase or Illumina small RNA adapter sequence was used. Prepare libraries for Illumina sequencing in less than a day. sequences, and is present in all adapters before the unique Index sequence. Illumina Advantage large-scale sequencing products feature lot-specific shipments and testing, extended shelf life, and advanced change notifications for greater laboratory efficiency. This product is also available as an Illumina Advantage (TG) product. The PCR reaction also adds index sequences on both ends of the DNA, thus enabling dual-indexed sequencing of pooled libraries on any. This change in base pair index codes requires adjustments to the sequencing run setup.įind robotic systems compatible with this kit Once hard-trimming of files is complete, Trim Galore will exit. FastQC does not indicate overrepresented sequences in the resulting FASTA-files, however still indicates Nextera Transposase Adapters in the 1 files. These unique dual index codes use 10 bp codes. SPECIFIC TRIMMING - without adapter/quality trimming -hardtrim5 Instead of performing adapter-/quality trimming, this option will simply hard-trim sequences to bp at the 5-end.The Illumina DNA UD Indexes Sets A, B, C, and D Indexes offer up to 384 unique dual indexing, which allows accurate assignment of reads and efficient use of the flow cell. Libraries prepared with Nextera XT kits are compatible with all Illumina RUO sequencers and Dx instruments in RUO mode only. Plus, the kit includes an innovative bead-based sample normalization that eliminates the need for library quantification prior to pooling and sequencing. At least, Nextera Transposase Sequence in FASTQC is defined as the reverse complement of the last 12 nucleotides of Read 1.Multiplexing of up to 384 samples per Nextera XT library is available for projects requiring greater throughput. In addition, Nextera adaptors are not in the default contaminant list of FASQC.Äoes anyone know if the above sequences are the corrected adapter Both analysis are quite different, and its possible to find adapter contamination in less than 0.1% of reads or in "readthrough" reads (so not over the full read length), and in both case, the contamination will not show up as overexpressed sequence. For more information on adapter trimming in Nextera Mate Pair, see the Data Processing of Nextera Mate Pair Reads on Illumina Sequencing Platforms technical note. In contrast, overexpressed sequences lists all of the sequence which make up more than 0.1% of the total over the first 50-75 nt. In cases where the circularized DNA is sheared close to the junction adapter, one read may again sequence into the adapter (Figure 2d). Yes, you should not necessarily expect that since the adapter (or more precisely, a 12-bp fragment of the adapter) is explicitly searched against your library. cause one read to sequence into the junction adapter downstream trimming of the junction adapter sequence will either remove this read entirely (Figure 2b) or else reduce the alignable length (Figure 2c). We performed deep sequencing of the Nextera-enriched dual-tagged library and of a. Would be picked up in overrepresented sequences section? or am i transposon ends (a) or transposon ends appended to unique adaptor sequences. One sequence is over represented: you have 306 reads which are exactly the sequence of the Nextera adapter. We will remove these duplicates later on. These sequences are provided for the sole purpose of understanding and publishing the results of your sequencing experiments. I expected the adapter sequence (Nextera Transposase Sequence) The read library quite often has PCR duplicates that are introduced simply by the PCR itself. Illumina Adapter Sequences This document provides the nucleotide sequences that comprise Illumina oligonucleotides used in Illumina sequencing technologies. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
